Simulate stochastic model
simstoch.RdSimulate a stochastic model generated with
uqsa::generateGillespieModel(), using the solver in this package.
Arguments
- ex
list of experiments, same as for the deterministic solvers.
- cmeModel
Either the cmeModel from as_cme, with a shared library path stored inside, or the path to the so file
- time.out
in seconds
- parameters
a numeric vector of appropriate size
Details
This will simulate all experimental conditions included in the list of experiments, including applying the inputs:
u <- experiments[[i]]$input - the input will be copied to the end of the model's internal parameter vector.
Like for deterministic models, we assume that there is a vector of
unknown parameter (a Markov chain variable, a vector of
optimization variables) and also known parameters (aka the input
parameters). The model itself does not distinguish between the two,
but one is the same between the experiments and one is different
between different experiments: modelParam <- c(mcmcParam, inputParam)
The path to the shared library, is required to contain at least one
slash in it, e.g.: "./model.so", "/tmp/Rsdkljhskjdhf/model.so" But,
not just "model.so", otherwise the shared library is interpreted as
a system library by dlopen() (it will not be found).
Examples
m <- model_from_tsv(uqsa_example("AKAR4"))
cme <- as_cme(m)
C <- generate_code(cme)
c_path(cme) <- write_c_code(C)
so_path(cme) <- shlib(cme)
#> Warning: running command 'PKG_CPPFLAGS='-O3 ' PKG_LIBS='-lgsl -lgslcblas -lm ' '/usr/lib/R/bin/R' CMD SHLIB /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c 2>&1' had status 1
#> using C compiler: ‘gcc (Ubuntu 13.3.0-6ubuntu2~24.04.1) 13.3.0’ gcc -I"/usr/share/R/include" -DNDEBUG -O3 -fpic -g -O2 -fno-omit-frame-pointer -mno-omit-leaf-frame-pointer -ffile-prefix-map=/build/r-base-FPSnzf/r-base-4.3.3=. -fstack-protector-strong -fstack-clash-protection -Wformat -Werror=format-security -fcf-protection -fdebug-prefix-map=/build/r-base-FPSnzf/r-base-4.3.3=/usr/src/r-base-4.3.3-2build2 -Wdate-time -D_FORTIFY_SOURCE=3 -c /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c -o /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.o /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c: In function ‘model_effects’: /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c:43:19: error: ‘_NA’ undeclared (first use in this function) 43 | a[_NA] = NA*NA; | ^~~ /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c:43:19: note: each undeclared identifier is reported only once for each function it appears in /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.c:43:26: error: ‘NA’ undeclared (first use in this function); did you mean ‘NAN’? 43 | a[_NA] = NA*NA; | ^~ | NAN make: *** [/usr/lib/R/etc/Makeconf:191: /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.o] Error 1
#> Warning: Building /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.so failed.
#> Warning: File /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.so does not exist
ex <- experiments(m)
p0 <- values(m$Parameter)
s <- simstoch(ex,cme)
#> Warning: File /tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.so does not exist (anymore).
#> Warning: The file «/tmp/RtmpgX4tEN/499b1cc1a9152200/AKAR4.so» does not exist.
res <- s(p0)
#> Error in s(p0): could not find function "s"
require(errors)
#> Loading required package: errors
plot(as.errors(ex[[1]]$outputTimes),ex[[1]]$data,xlab="time",ylab="AKAR4p",main=names(ex)[1])
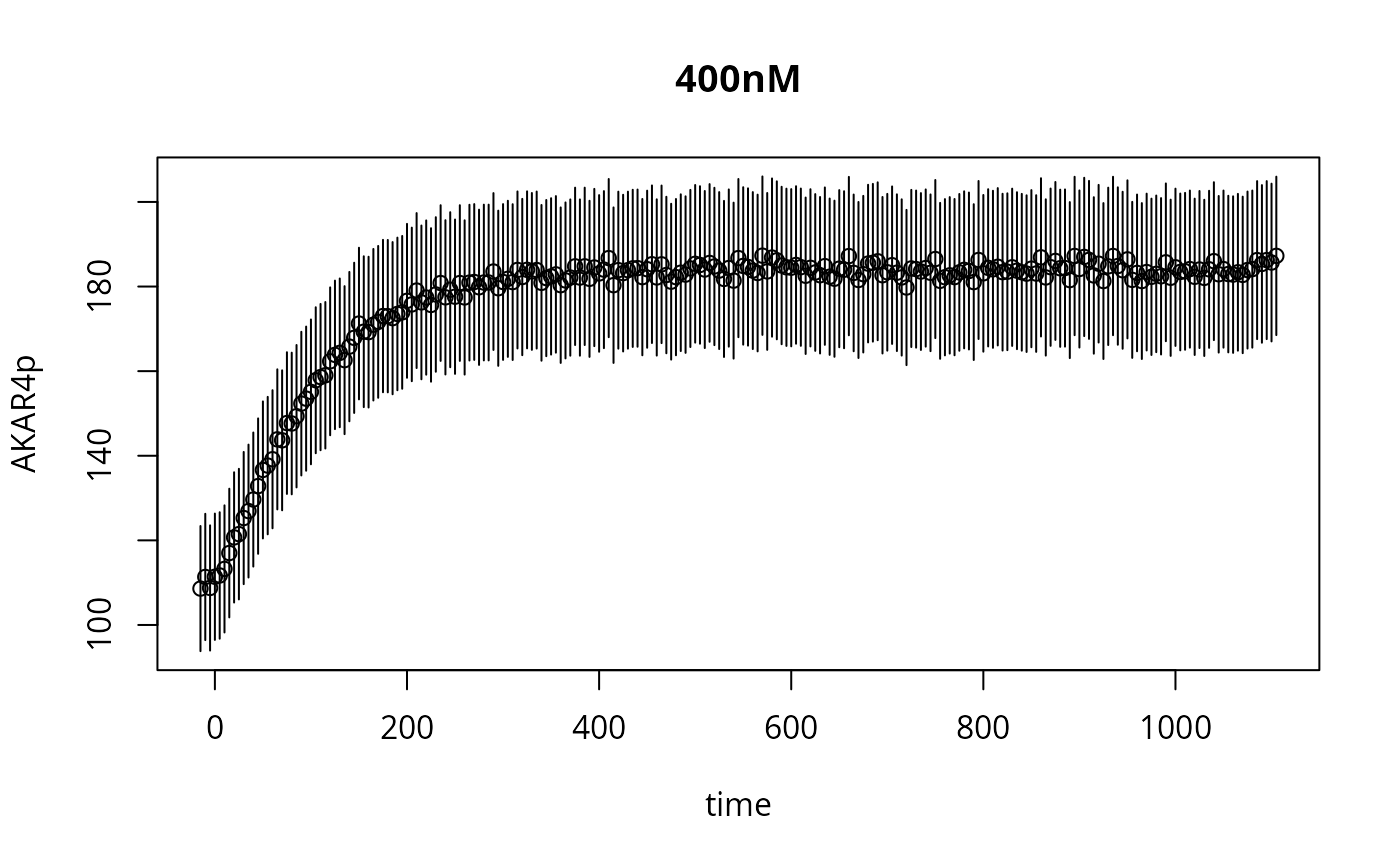 lines(ex[[1]]$outputTimes,res[[1]]$func,type="s",lwd=2,col="red3")
#> Error in eval(expr, envir, enclos): object 'res' not found
lines(ex[[1]]$outputTimes,res[[1]]$func,type="s",lwd=2,col="red3")
#> Error in eval(expr, envir, enclos): object 'res' not found