Uncertainty Quantification of AKAP79 (deterministic model)
sampleAKAP79.Rmd
library(uqsa)
library(parallel)
options(mc.cores=detectCores())To take a sufficiently large sample of AKAP79, we shall use MPI and
parallel tempering. There are several ways to launch MPI processes
(e.g. spawn processes). We will use the simplest form: launch all MPI
workers right from the start, using mpirun -N 8 (or
similar) and have all processes communicate on
MPI_COMM_WORLD, this requires very minimal MPI code in R.
This approach doesn’t allow us to adapt the number of temperatures,
which is probably not optimal but makes it easier to write and
understand the code and increases performance significantly compared to
a sequential Markov chain.
The R code we are going to execute is not a code block on this page
that we run in an interactive R session. Instead, it has to be a file
that we pass on to mpirun.
In this article we show the results of a parallel tempering SMMALA (simplified manifold Metropolis adjusted Langevin algorithm) run. The article titled Parallel chains with MPI explains how to prepare the R-script for mpirun execution – it includes the listing of the script we used in this article.
We will store all files related to this in a very predictable
location: /dev/shm; not R’s tempdir(). This is
because the R blocks in this article are not executed in the same R
session as the MPI workers will run in. Each MPI worker is its own
process; thus they all have different temporary directories. The
directory /dev/shm is a virtual directory that lives in the
machines RAM. It is standard on GNU/Linux systems, but not on MACOS, so
this example will not work verbatim there. The sampling will take a
while, even for a very small sample, so it is not recommended to try out
this article.
Create C Source Code
First we load the model in its SBtab form and create all derived model files. We do this step only once and have the MPI part of the process load the coimpleted results of this:
f <- uqsa::uqsa_example("AKAP79")
m <- model_from_tsv(f)Create the ODE interpretation of m:
o <- as_ode(m)
#> Loading required namespace: pracma
cat(generate_code(o),sep="\n",file="/dev/shm/AKAP79.c")
c_path(o) <- "/dev/shm/AKAP79.c"
so_path(o) <- shlib(o)
saveRDS(o,file="/dev/shm/AKAP79-ode.RDS")Load the simulation instructions for o:
ex <- experiments(m,o)
#> Warning in experiments(m, o): CONFLICT: This model seems to have event based
#> transformations and conservation laws. These two concepts clash with one
#> another if a compound is conserved, but also changed by scheduled events.
saveRDS(ex,file="/dev/shm/AKAP79-ex.RDS")There is a high-level function that does most of what we need to do automatically:
p <- values(m$Parameter)
smmala <- high_level_smmala(m,o,ex,p)
#> The parameters are given in log10-scale, so the simulator will do the reverse transformation: 10^p.Plain Sample
h <- tune_step_size(smmala)
#> acceptance rate: 0.02, step-size: 0.0001;
#> acceptance rate: 0.06, step-size: 1.27186e-06;
#> acceptance rate: 0.37, step-size: 1.38538e-07;
#> acceptance rate: 0.05, step-size: 1.9023e-07;
#> acceptance rate: 0.2, step-size: 1.46331e-08;
#> acceptance rate: 0.19, step-size: 1.14209e-08;
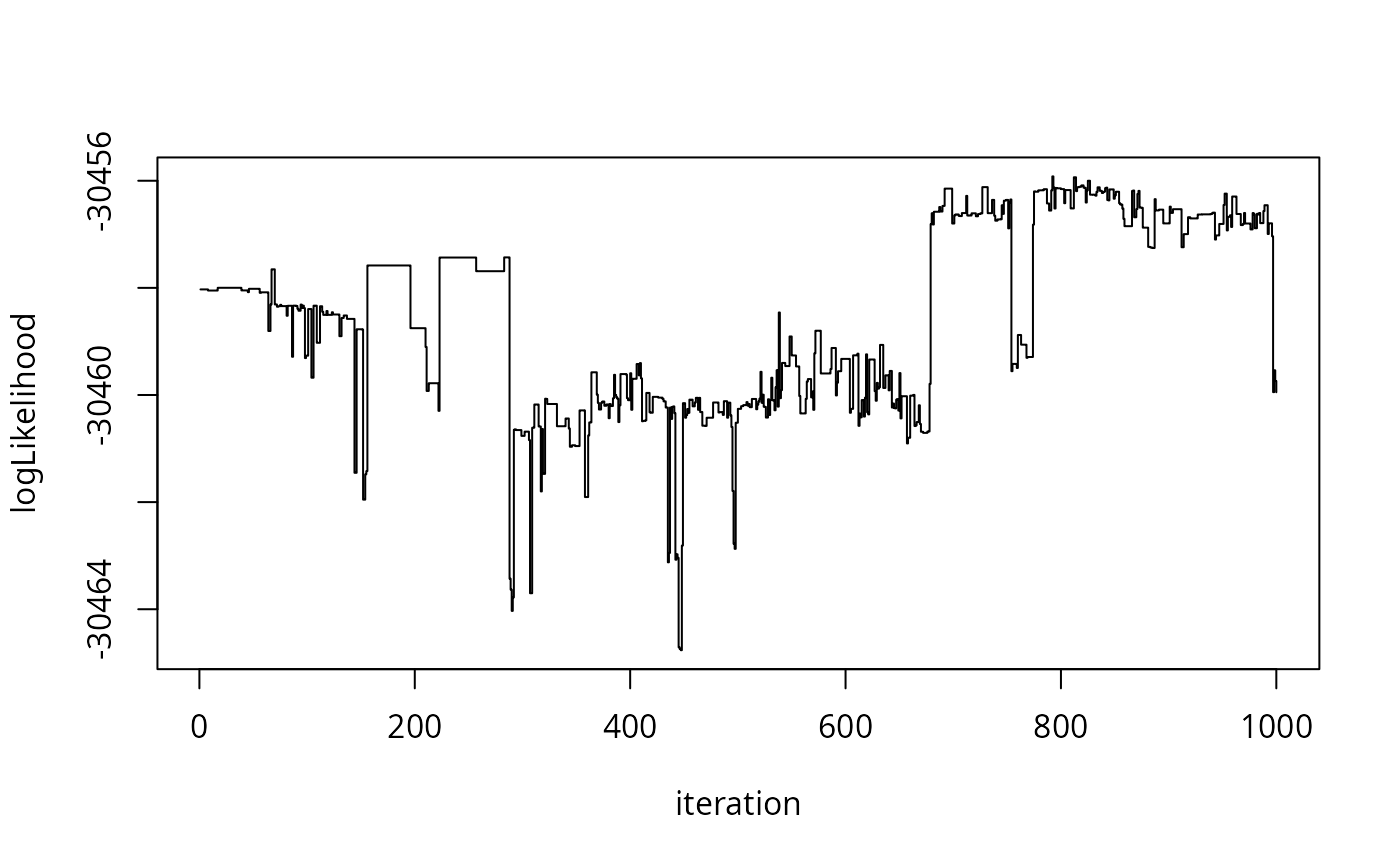
X <- smmala(smmala %@% "init",1000,h)
plot(
X %@% "logLikelihood",
type="s",
xlab="iteration",
ylab="logLikelihood"
)
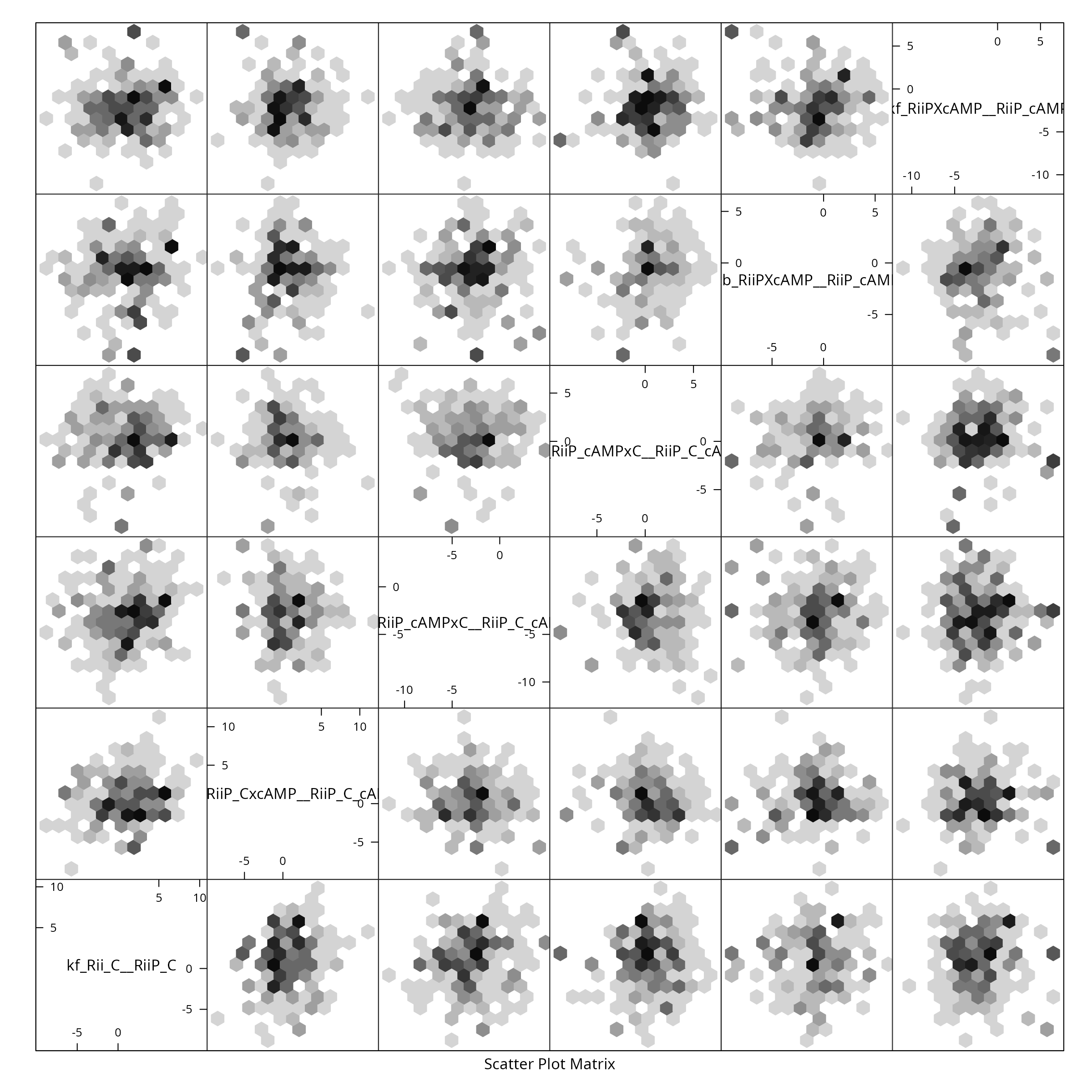
We shall only plot the pairs of the first 6 dimensions (this is a big model):
if (require(hexbin)){
hexplom(X[,seq(6)])
} else {
pairs(X[,seq(6)])
}
#> Loading required package: hexbin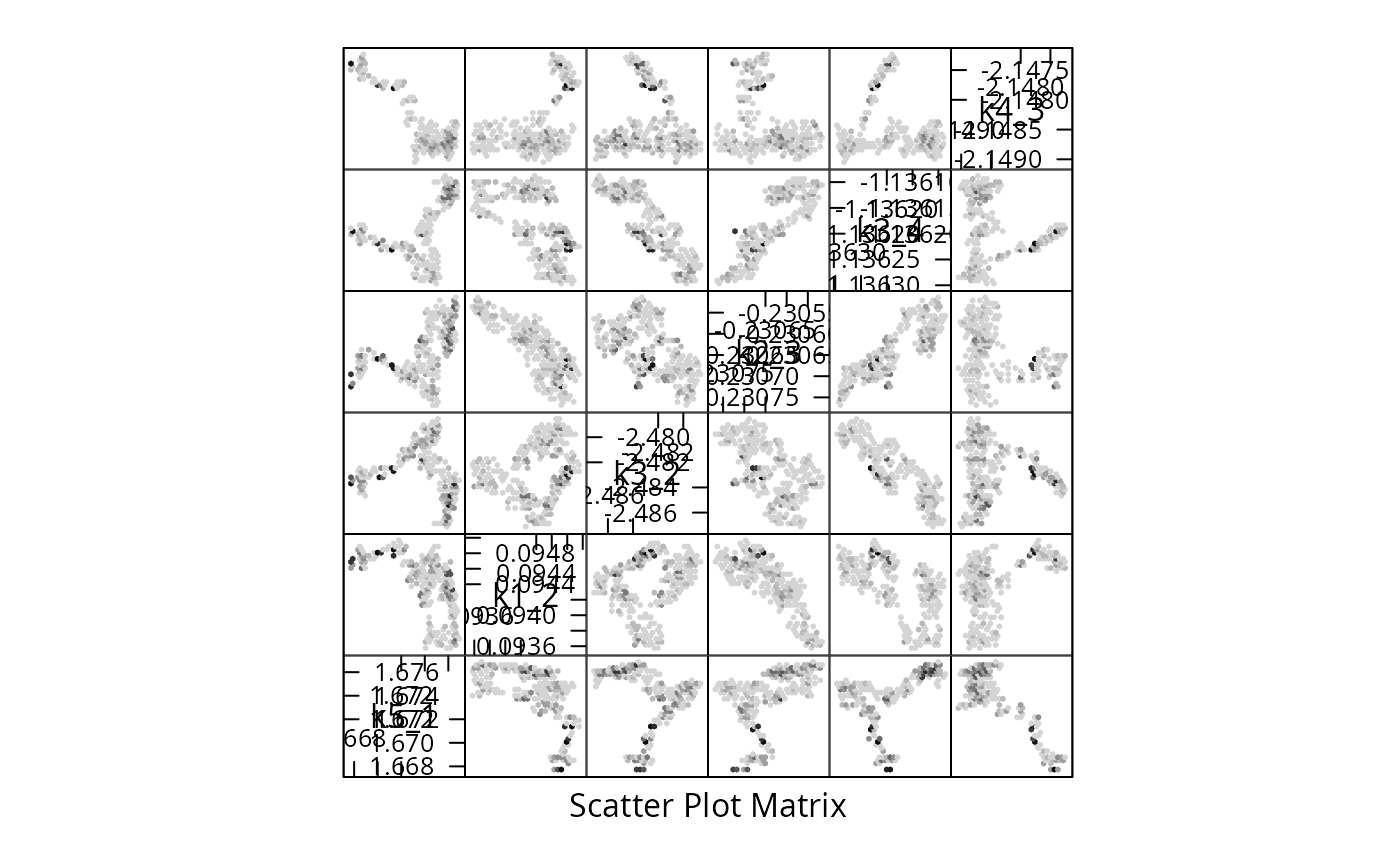
Sample via MPI
Here we run a prepared R script from the command line (any POSIX shell will do: bash/zsh/fish/dash)
N=4 # default number of MPI workers
[ -e '/proc/cpuinfo' ] && N=$((`grep -c processor /proc/cpuinfo`)) && nm="`grep -m1 'model name' /proc/cpuinfo`"
echo "We will use $N cores with $nm"
start_time=$((`date +%s`))
date
mpirun -H localhost:$N ./pt-smmala-akap79.R 10000 > /dev/null 2>&1
date
end_time=$((`date +%s`))
echo "Time spent sampling: $((end_time - start_time)) seconds ($(( (end_time - start_time)/60 )) minutes)."
#> We will use 8 cores with model name : Intel(R) Core(TM) i7-7700HQ CPU @ 2.80GHz
#> Mon 04 May 2026 06:13:51 PM CEST
#> Mon 04 May 2026 07:32:23 PM CEST
#> Time spent sampling: 4712 seconds (78 minutes).On a high performance computing (hpc) cluster, the above would be in a slurm script or similar workload manager.
Inspect the Results
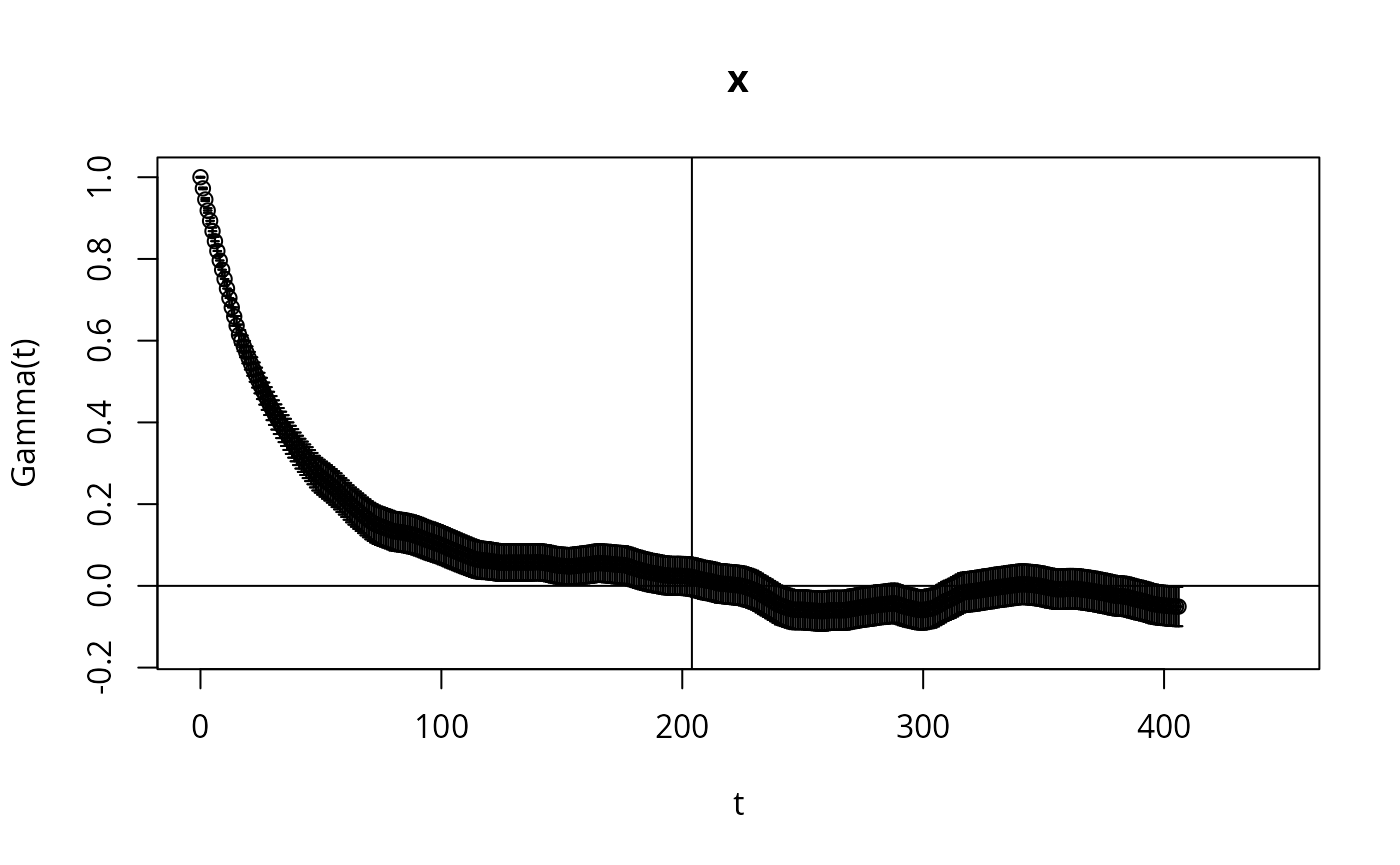
Here we check the integrated auto-correlation length (Markov chain time)
Sample <- readRDS(file="AKAP79-temperature-ordered-pt-smmala-sample-2-for-rank-0.RDS")
n <- NROW(Sample)
l <- attr(Sample,"logLikelihood")
plot(l,type="l",xlab="mcmc index",ylab="log-likelihood")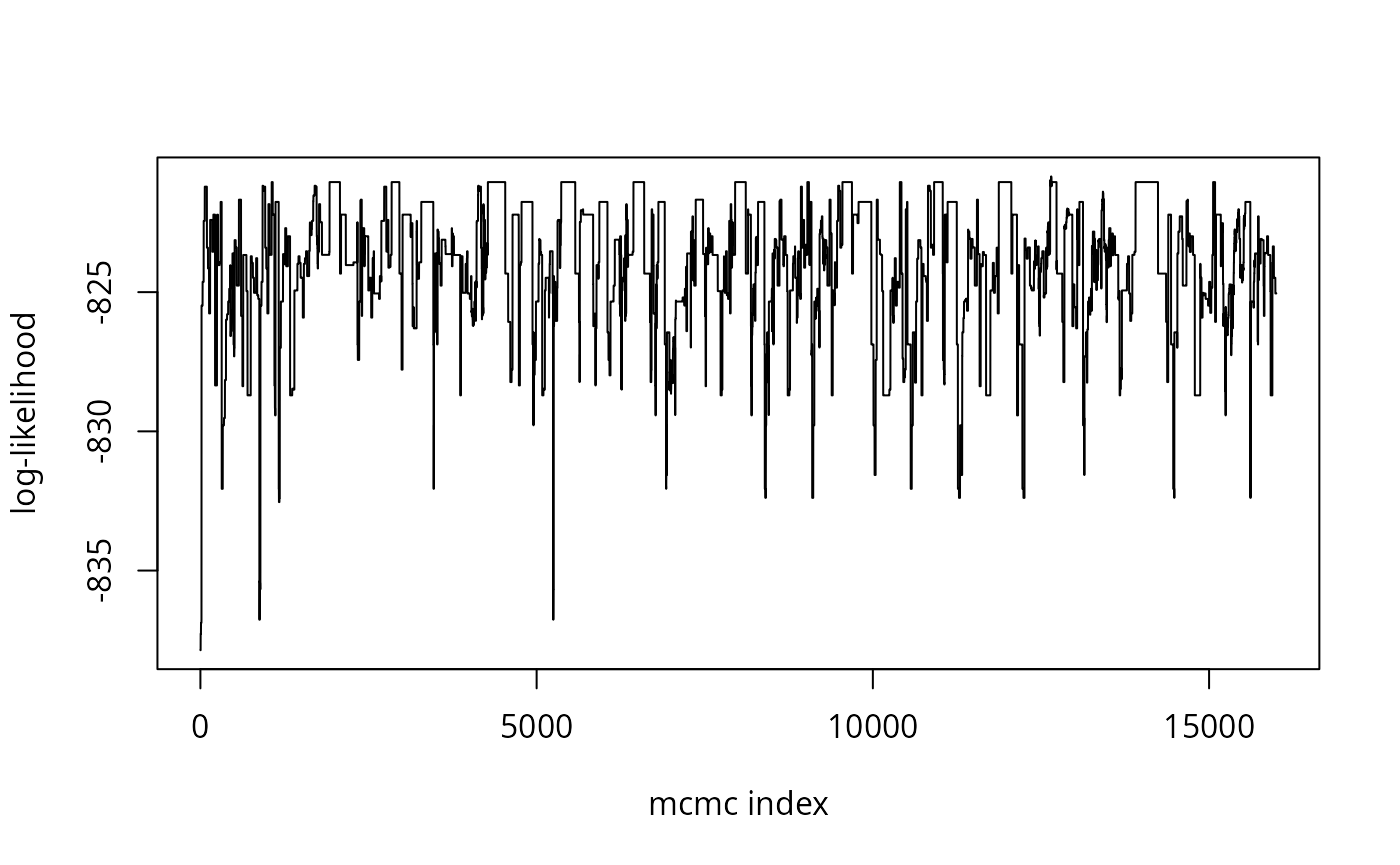
if (require(hadron)){
res <- hadron::uwerr(data=l,pl=TRUE)
tau <- ceiling(res$tauint + res$dtauint) # upper bound
} else {
A <- acf(l,lag.max=10000)
tau <- ceiling(sum(A$acf))
}
#> Loading required package: hadron
#>
#> Attaching package: 'hadron'
#> The following object is masked from 'package:base':
#>
#> kappa
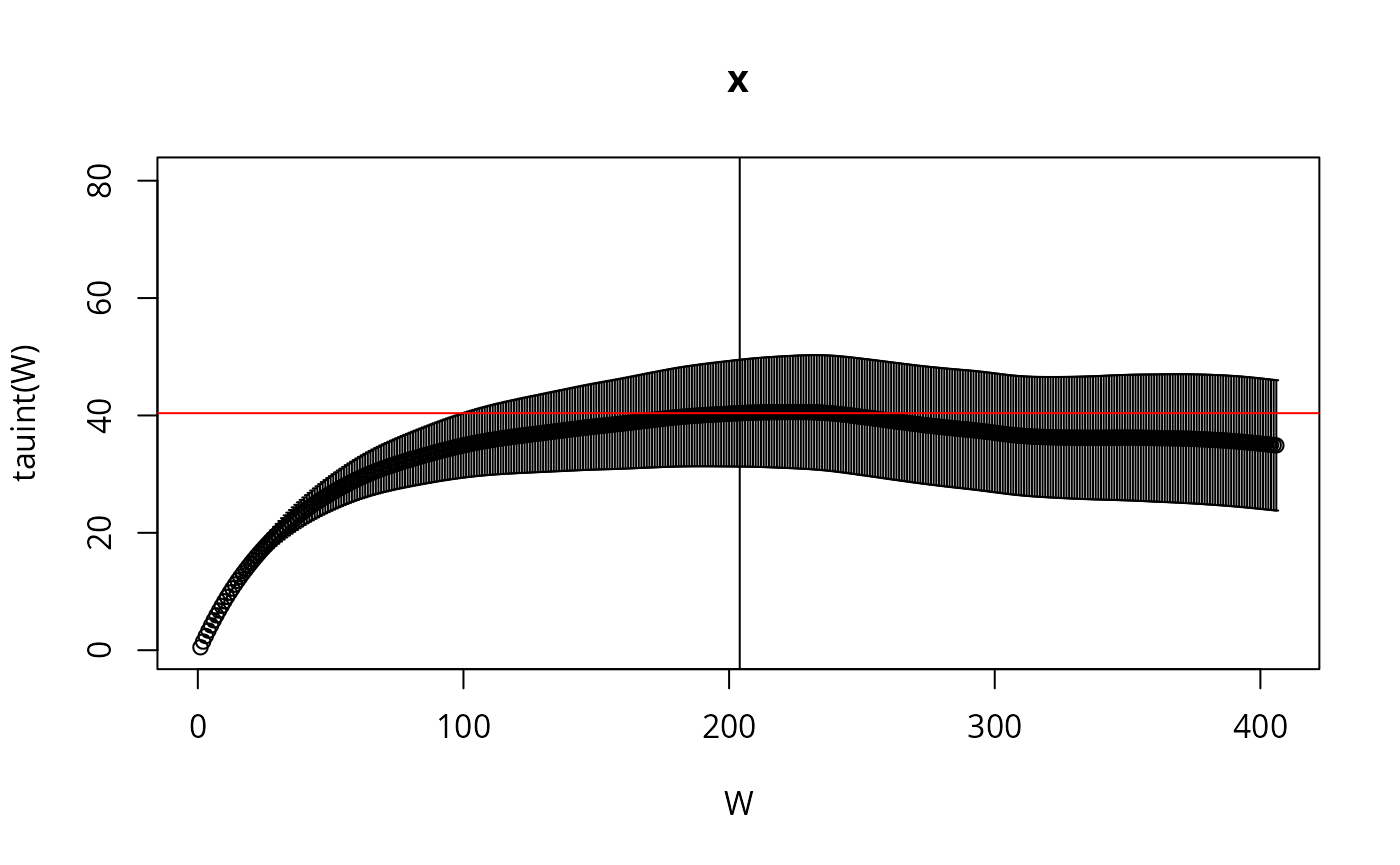

We reduce the sample for plotting purposes, by showing only every \(\tau_{\text{int.}}\)-th point and create a pairs plot for the first 6 parameters:
X <- Sample[seq(1,n,by=tau),]
colnames(X) <- rownames(m$Parameter)
if (require(hexbin)){
hexplom(X[,seq(6)],xbins=12)
} else {
pairs(X[,seq(6)])
}