Global sensitivity analysis on AKAR4 - independent input factors
GSA_AKAR4.RmdThis article provides code do perform global sensitivity analysis with the Sobol-Saltelli method and the with the binning-approach.
Load the model files, create ODE model code and load examples similar to the example Simulate the AKAR4 deterministic model.
f <- uqsa_example("AKAR4")
m <- model_from_tsv(f)
o <- as_ode(m,cla=FALSE)
C <- generate_code(o)
c_path(o) <- write_c_code(C)
so_path(o) <- shlib(o)
ex <- experiments(m,o)Create simulator
We construct a simulator function and and test it on default parameter values.
options(mc.cores = length(ex))
s <- simulator.c(ex,o,parMap=log10ParMap,omit=3)
p <- log10(values(m$Parameter))
y <- s(p)Set meta-parameters for the global sensitivity simulation
nSamples <- 40000nSamples corresponds to the number of samples in M1 (or M2) of the Soboll-Saltelli approach. The number of simulations of the Sobol-Saltelli approach consists of 2*nSampels+nPars*nSamples number of simulations. In the binning approach below we use 2*nSamples number of samples (corresponding to 2*nSamples number of simulations) to use the same number of independent sample points as Sobol-Saltelli.
Construct parameter prior samples according to Sobol-Saltelli (M1, M2, N).
rprior <- rNormalPrior(p, array(1, length(p)))
prior <- saltelli_prior(nSamples, rprior)
names(prior)
#> [1] "M1" "M2" "N"Plot parameter prior (M1)
title <- paste("Prior distribution experiment");
boxplot(prior$M1, main = title, names=names(p), las=2, cex.main=0.9, cex.axis=0.3)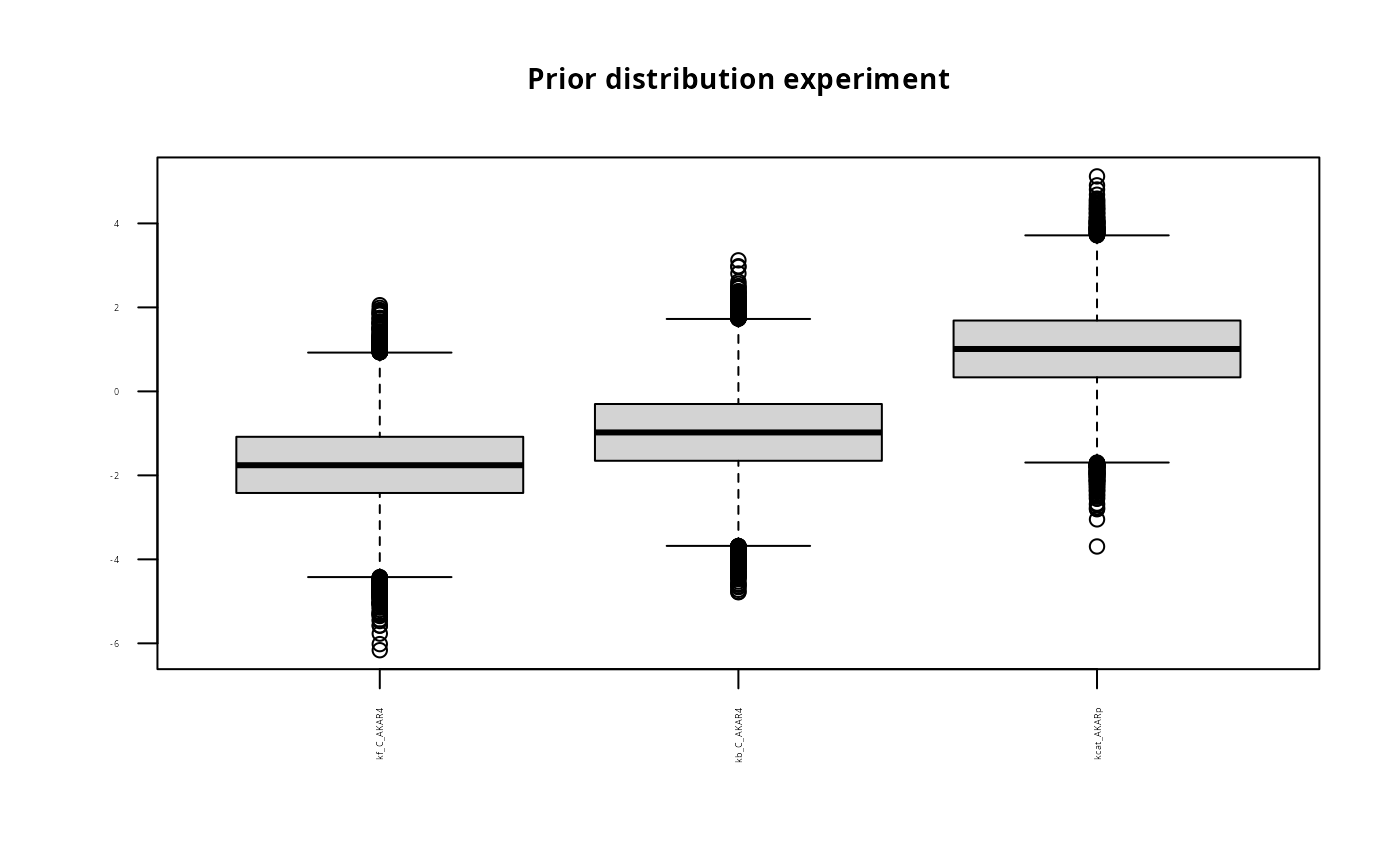
Calculate and plot sensitivity indexes
Calculate sensitivity indexes for sobol-saltelli
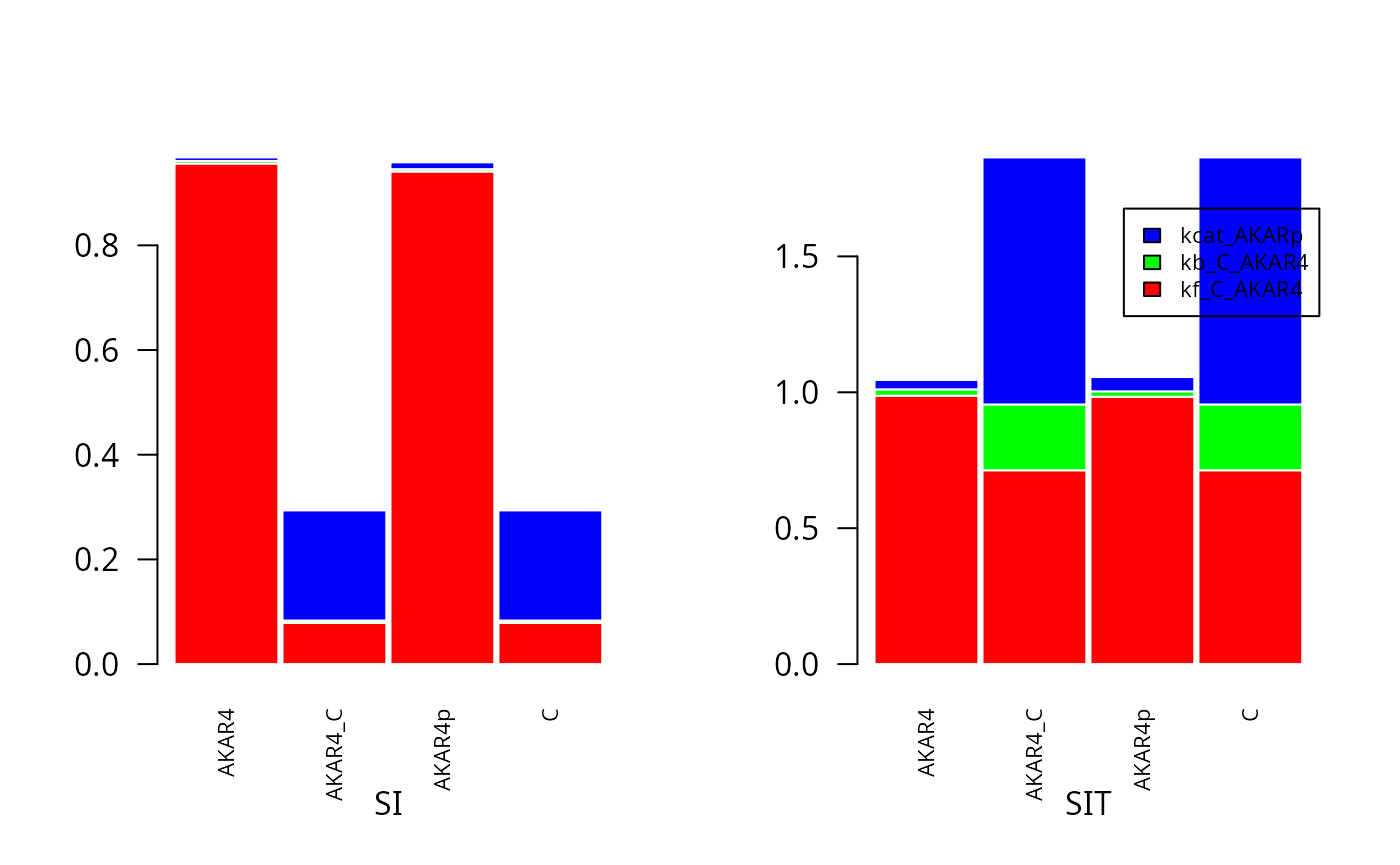
SA <- gsa_saltelli(fM1,fM2,fN)Plot first (SI) and total-order (SIT) sensitivity indexes for sobol-saltelli
cols=rainbow(3)
par(mfrow = c(1, 2))
barplot(
t(SA$SI[,]),
col=cols,
border="white",
space=0.04,
cex.axis=1,
names.arg=names(ex[[E]]$initialState),
cex.names = 0.7,
las = 2,
xlab="SI",
las=2,
cex.main=0.9
)
barplot(
t(SA$SIT[,]),
col=cols,
border="white",
space=0.04,
cex.axis=1,
names.arg=names(ex[[E]]$initialState),
cex.names = 0.7,
las = 2,
xlab="SIT",
legend.text=names(p),
args.legend = list(x = "topright", inset=c(0, 0.1), cex=0.7)
)
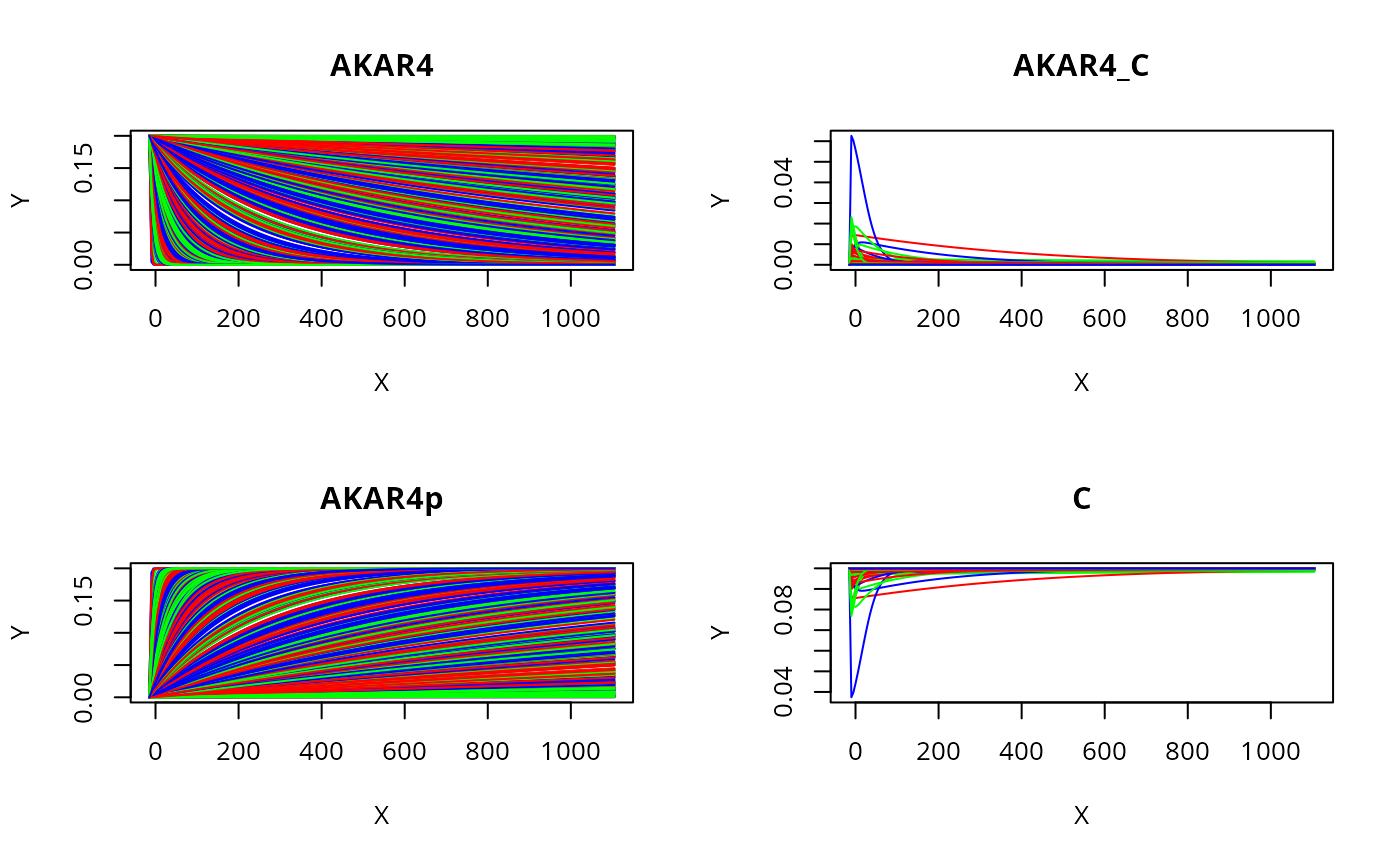
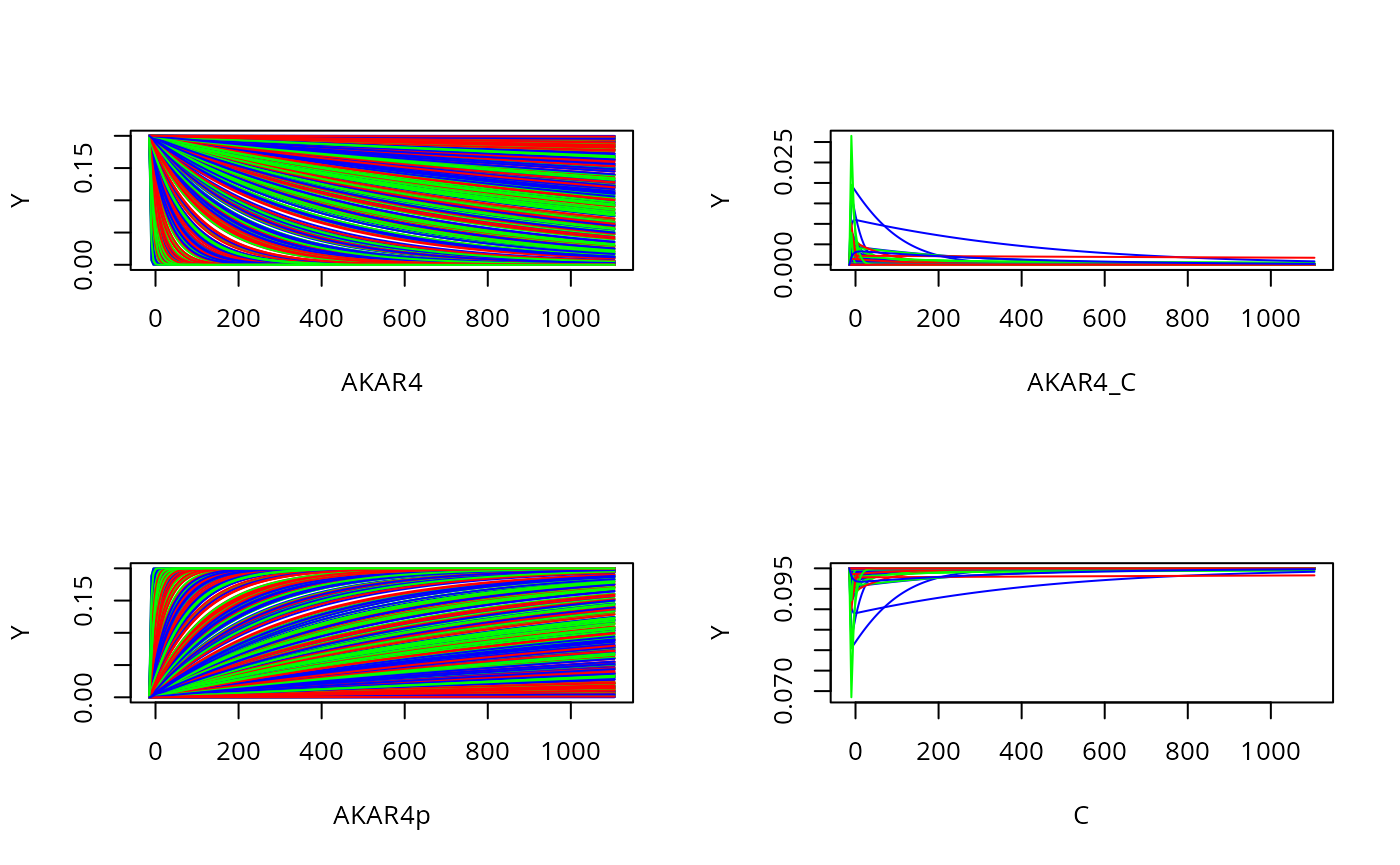
Plot all states and timeponts for the experiment
allTimesSample=s(t(prior$M2))[[E]]$state
par(mfrow = c(2, 2))
for (i in seq_along(ex[[E]]$initialState)) {
matplot(
ex[[E]]$outputTime,
allTimesSample[i,,seq(500)],
type = "l",
lty = 1,
col = c("red", "blue", "green"),
ylab = "Y",
xlab = names(ex[[E]]$initialState)[i]
)
}
Calculate first order sensitivity index (SI) based on binning approach
SIappr <- gsa_binning(rbind(prior$M1,prior$M2), rbind(fM1,fM2))Plot SI for Sobol-Saltelli versus the binning approach
par(mfrow = c(1, 3))
barplot(
t(SA$SI[,]),
col=cols,
border="white",
space=0.04,
cex.axis=1,
names.arg=names(ex[[E]]$initialState),
cex.names = 0.7,
las = 2,
main="SI Saltelli")
barplot(
t(SIappr),
col=cols,
border="white",
space=0.04,
cex.axis=1,
cex.names = 0.7,
las = 2,
main="approximate SI")
barplot(
c(0),
axes=FALSE,
col=cols,
border="white",
space=0.04,
font.axis=2,
legend.text=names(p)
)
mtext(paste("Comparision GSA methods, sample size=",as.character(2*nSamples)), side = 3, line = -1.2, outer = TRUE)