Transformation Events
events.RmdThe solvers we use in most of our examples have the built-in ability to perform sudden interventions. Here we show an example of how this can be used to trigger sudden spikes (action potentials) in a neuroscience context.
f <- uqsa_example('Spike')
m <- model_from_tsv(f)
o <- as_ode(m,cla=FALSE) # no conservation law analysis
ex <- experiments(m,o)
C <- generate_code(o)
c_path(o) <- write_c_code(C)
#> Writing file: /tmp/RtmpGHPWIK/b9ad1a93e783d1c5/Spike.c
so_path(o) <- shlib(o)The transformations are still present in m:
m$Transformation
#> Ca Buffer
#> APCa Ca A*(k2-k1)*doseBut, also built into the model and simulation instructions
ex
ex[[1]]$events
#> $time
#> [1] 0 50 100 150 200 250 300 350 400
#>
#> $label
#> [1] 0 0 0 0 0 0 0 0 0
#>
#> $dose
#> [1] 1 1 1 1 1 1 1 1 1Here, an event has a label, which is just the c-style offset
into the Transformation table. Offset 0 specifies the first
transformation. Each line of the transformation table is a code block
inside of a C switch statement. So, the label also selects the case of
that switch statement, see further below.
Each event has a scalar dose variable, currently this
lower-case dose variable cannot be renamed. It specifies
the intensity of the transformation. If the transformation requires more
than one parameter they have to be defined explicitly as part of the
model’s state, or input (Input table).
Because events can change Input values, we can also define an input that becomes a sort of storage unit that can remember the value of a variable at a given time. So, we could define an output function that uses this time-spcificity of events to define more intricate return values.
Events can change state variables, but also the inputs. They can also
depend on the time t.
Simulation
tm <- seq(-4,1000)
ex[[1]]$outputTimes <- tm
SIM <- simulator.c(ex,o)
par <- values(m$Parameter)
print(par)
#> k1 k2 A
#> 9.78422e-03 3.44800e-02 2.29127e+02
#> attr(,"unit")
#> k1 k2 A
#> "1/s" "1/s" "1/s"
res <- SIM(par)
y <- as.numeric(res[[1]]$func)
plot(tm,y,bty="n",type='l',xlab="t",ylab="Ca")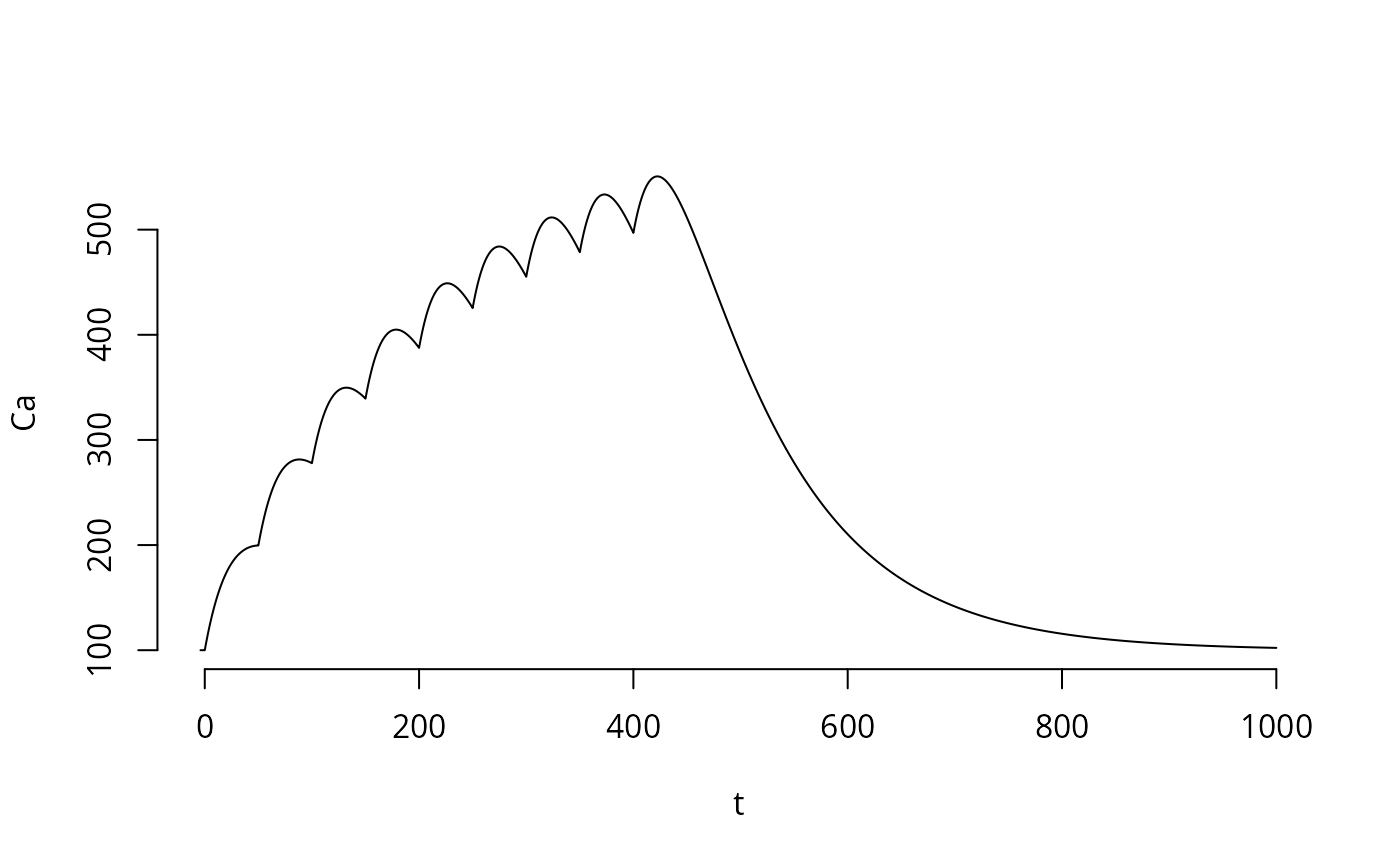
One Calcium Spike
Maybe, we should also simulate how one Ca spike would look like, and also try reducing the dose a bit:
ex[[1]]$event$time <- 0.0
ex[[1]]$event$label <- 0
ex[[1]]$event$dose <- 1.0
ex[[2]] <- ex[[1]]
ex[[2]]$event$dose <- 0.6
SIM <- simulator.c(ex,o)
res <- SIM(par)
y1 <- as.numeric(res[[1]]$func)
y2 <- as.numeric(res[[2]]$func)
plot(tm,y1,bty="n",type='l',xlab="t",ylab="Ca")
lines(tm,y2,lty=2)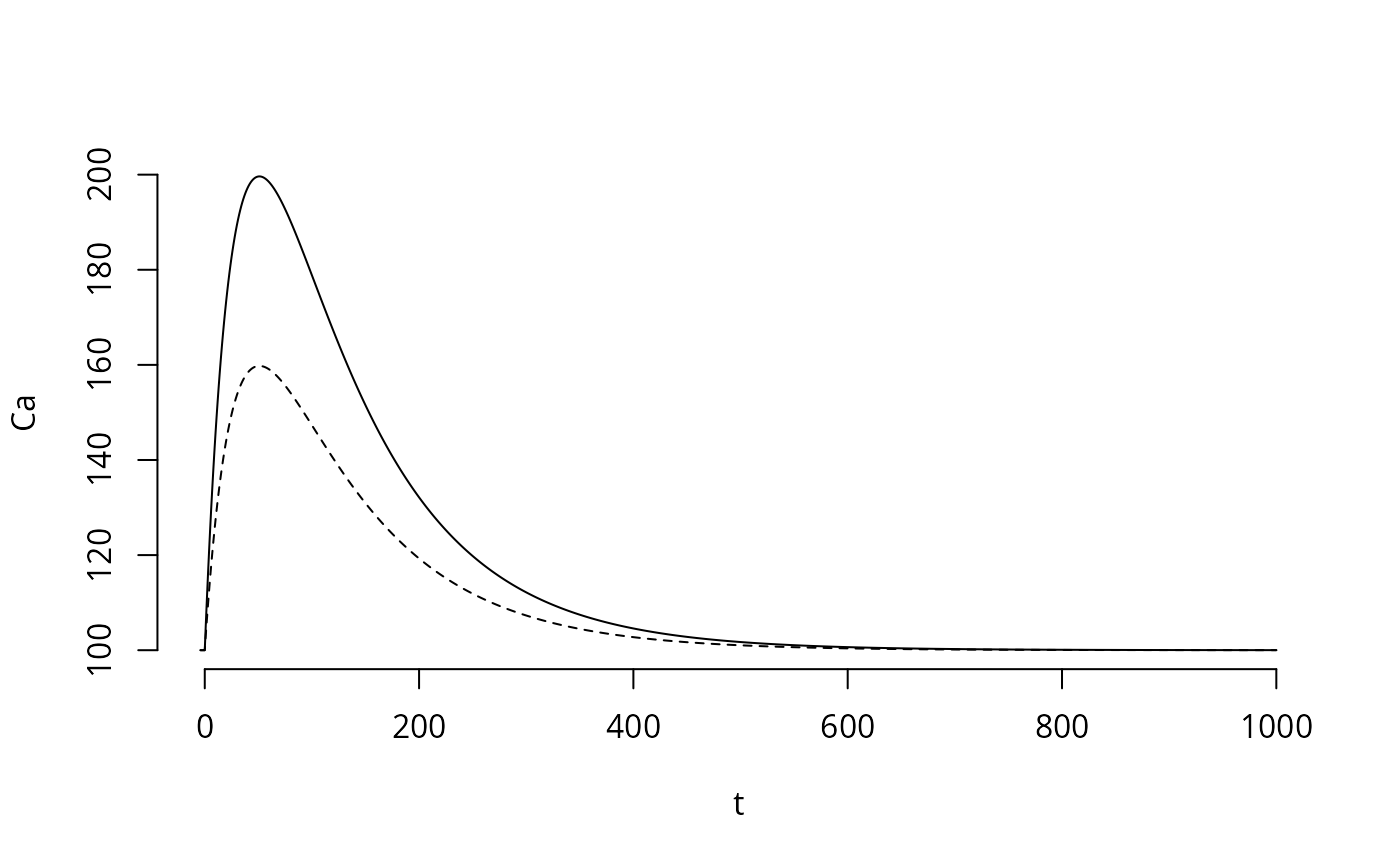
Details
In this section, we provide details about the implementation of scheduled events. This information could be useful in writing complex simulation instructions. Some concepts from systems biology cannot be expressed in the model files as they have a very plain set of rules. But with some knowledge of the event system, we can express fairly intricate simulation instructions.
Switch Statement
This specific model has only one available transformation, the only
valid label is 0 and corresponds to the enum
value APCa.
/* ... */
enum eventLabel { APCa, numEvents };
/* ... */
/* skipped functions */
/* ... */
int Spike_event(double t, double *y_, double *p_, int EventLabel, double dose){
if (!y_ || EventLabel<0) return numEvents;
double k1 = p_[_k1]; /* [ 0] */
double k2 = p_[_k2]; /* [ 1] */
double A = p_[_A]; /* [ 2] */
double CaBase = p_[_CaBase]; /* [ 3] */
double Ca = y_[_Ca]; /* [ 0] */
double Buffer = y_[_Buffer]; /* [ 1] */
double rf1 = Buffer;
double rf2 = -k1*k2*Ca-(k1+k2)*Buffer;
switch(EventLabel){
case APCa:
y_[_Buffer] = A*(k2-k1)*dose;
break;
}
return GSL_SUCCESS;
}The C cource defines enums for the event labels to make the code more
readable and identify the transformations (case labels) by
name.
The transformation expression A*(k2-k1)*dose has access
to all constants, parameters (input or otherwise), and expressions such
as reaction fluxes.
This function is called when an event is scheduled to occur. The integrator is then reset as if a new integration were started from the changed model state (this does not reset the input values). This is different from triggered events, which happen when a condition is met. Triggered events are not implemented but exist in other packages/toolboxes/languages such as COPASI.
Inputs as Read/Write Buffers
Systems Biology often requires fairly difficult to express
experimental setups. Relative Data (Experiment and Control) are phrased
in such a way that two simulations are performed for each data (pair of)
meaningful data. If the experiment measures \(A\) and \(\prime
A\) where \(\prime A\) is the
control, and their ratio \(a\)
is meaningful, we should compare that ratio after simulating the
scenarios for A and then for A'.
This could be either two experiments in the Experiments table and a custom likelihood function that knows to take the ratio. Or, a simulation that receives to different parameter vectors (as a two-column-matrix). And defines a transformation event that stores a value in an input (which retains its value) for the next parameter column.
The experiment list ex will not have its values changed
of course. This retention happens in an internal buffer that is
read/write accessible to the ODE solver. The final state of the
input-vector it uses is not returned in any way.
Each experiment in the list is assigned its own input buffer, which is re-used when the next parameter vector is processed, this is actually true for the input parameters and the model’s kinetic parameters as the ODE solver doesn’t distinguish these two. The solver works with a matrix of size \(N\times P\) where \(N\) is the number of experiments and \(P\) is the number of overall parameters. The parameters are modified by enum label, so there is no need to know where in the array they are stored.